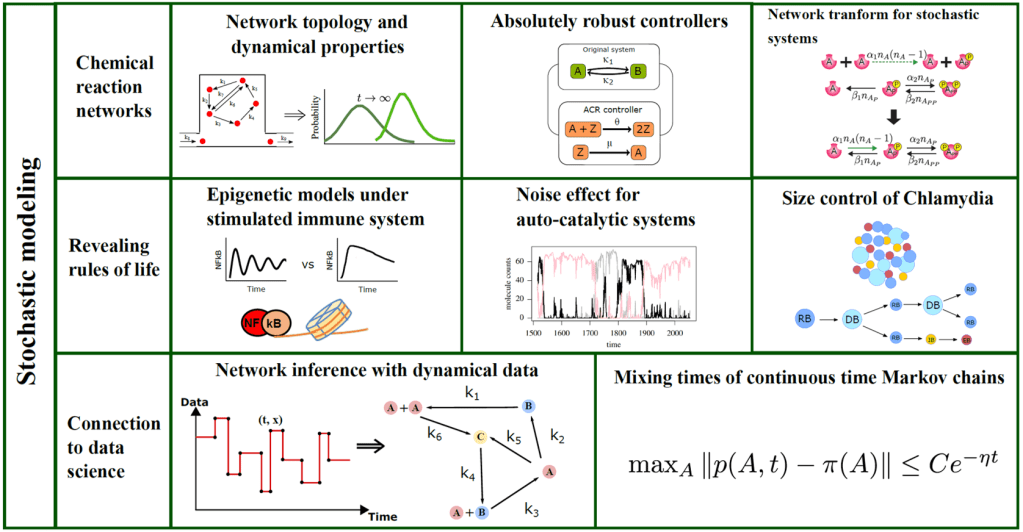
Stochastically modeled reaction networks: A biochemical system is described with a graphical configuration called a reaction network. These graphical configurations are used to model broad classes of interaction systems including signaling systems, viral infections, metabolism, neuronal networks, population models, etc. One of the most challenging issues systems biologists face is the extraordinarily complicated structure of a system. Thus, characterizing the system structure is a major open question in the field as the structure induces emergent phenotypes and behaviors of the system’s dynamics.

To find other network characterization, my colleges and I have used a hierarchical structure of given reactions, so-called `generalized tier-structures’, to discover new network conditions guaranteeing that the associated stochastic process admits a stationary distribution. Those network conditions can be satisfied in more realistic models such as i) fully-open bimolecular systems with a single weakly reversible component, ii) bimolecular system with dimerization of each species, or iii) bimolecular system with a single linkage class including all the dimers or monomers [1, 4, 5]. Importantly, the stability holds regardless of the choice of system parameters for the model, which are typically unknown in practice.
Absolutely robust controllers: By using the long-term stability of reaction systems, my colleague and I constructed absolutely robust controllers that can control the distribution of molecular counts in a given stochastic system. As opposed to controlling deterministic models, we have additional control goals for stochastic models because it is important to control not only the mean expression level but also its variance (i.e. noise) and ideally the full probability distribution of the target species. In [3], we showed that if some stability condition is met, the target species in the controlled system follows a Poisson distribution centered at a user-defined value. Since the control species can suffer from complete extinction, we set high initial abundance for the control species. As the control species has the higher order of copy numbers, we have used multi-scaling analysis to prove the long-term stability of the controlled system and in turn verified controllability of our synthetic circuit.

Not only the stochastic models but also the target species in a deterministic model can be controlled with our controller as its concentration is driven to the desired steady state value. We also have emphasized that the controlled system under both deterministic and stochastic settings is resistant to external disturbances such as parameter changes and additional in- and out-flows. This property is known as robust perfect adaptation, which is a highly desirable goal in control theory.
Computing rare event probabilities via network modifications: We are often interested in rare events of a stochastic model such as the extinction of diseases. Stochastic simulation algorithms including Gillespie’s algorithm have been typically employed to estimate the probability distribution of a Markov process. However, when the target event rarely occurs, significantly long simulations may be required to sample enough trajectories to estimate the distribution of the system. Alternatively, the chemical master equation, which is a system of ordinary differential equations, can be utilized to compute the rare event probability. However, it can suffer from dimensionality as a stochastic system often admits infinitely many states.
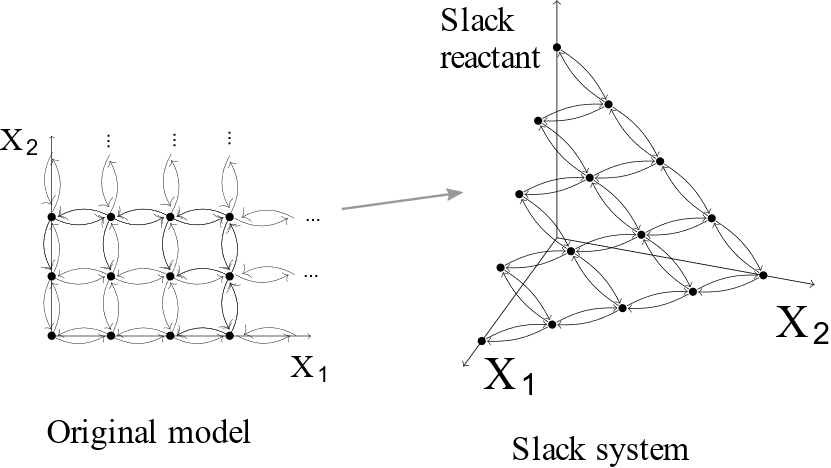
In [8], we proposed a new network modification method that automatically truncates the state-space. The modified network composed of `slack reactants’ added to a given network has approximately the same dynamic as the original system. We proved that the modified system converges to the original system as the size of truncation increases. We also provided the application of our theoretical frameworks with practical reaction network models.
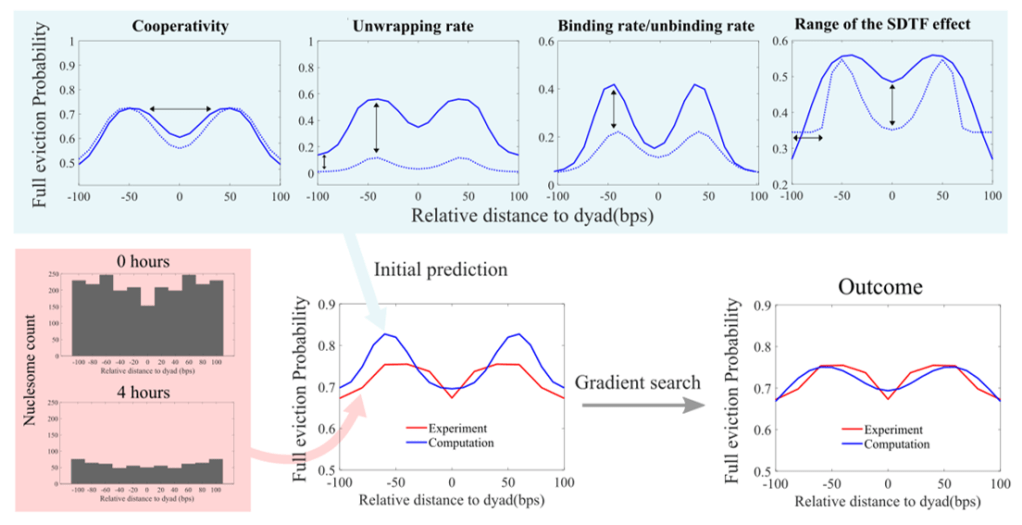
Evidence of cooperativity in in vivo immune cells: I collaborated with biologists in Alexander Hoffmann’s lab to investigate mechanistic principles and biophysical parameters that allow SDTF binding to generate distinct epigenomes in living organs, in the context of pathogen responses. By utilizing experimental measurements of location-specific nucleosome counts with ATAC-seq, a next-generation sequencing technology, we detected from the experimental data evidence of cooperativity in nucleosome unwrapping steps. We then investigated the influence of SDTF-binding location on nucleosome eviction and estimated the system parameters using reverse engineering. Our efforts provide rules governing nucleosome dynamics in vivo that can predict how nucleosomes respond to SDTF activation at specific genomic locations to produce epigenomic alterations during inflammation.
Size control mechanism of chlamydia in a host cell: I am currently supported by an NIH research grant for researching the size-control mechanism of chlamydia. Chlamydia is a bacteria causing widespread human diseases by infecting eukaryotic host cells. Unlike the well-known cell division mechanism that typically maintains the size of each cell, the average size of chlamydia decreases during its developmental cycle. The size-control mechanism was proposed as the replicating forms (RB) become progressively smaller through repeated cycles of replication and convert into the infectious forms (EB) only below a size threshold (Lee et. al. 2018). Our current project is to test our hypothesis experimentally and mathematically. We use a branching process to describe division and conversion of chlamydia and to study the size-control mechanism.
Effect of intrinsic noise in biological systems: A stochastic system is not necessarily understood as a process fluctuating around the trajectory of the deterministic counterpart. Many examples are known to have a discrepancy between stochastic and deterministic modeling. For instance, cell polarity can be induced by oscillations in the stochastic system, while it is not found for comparable parameters in the deterministic case [2]. Moreover, a stochastic model can admit multi-peaks in the long-term distribution, which can represent multiple potential phenotypes in a single cell system, while the deterministic counterpart admits only a single steady state. Although such an effect of intrinsic noise was proposed in many systems, a rigorous verification was not often provided. In [7], we proved stability and finite moment conditions for a class of autocatalytic systems. We also verified the effect of the intrinsic noise in the system by finding that the stationary distribution of the autocatalytic systems is Dirichlet-multinomial distribution, which has multi-peaks when the system volume is small. This multimodality is not realized in the deterministic counterpart that has a single stable steady state.

Biochemical network inference: The structure of biochemical networks and the associated parameters are typically unknown in practice. Many theoretical and computational methods such as Bayesian inference and machine-learning tools were proposed to infer the system parameters and the underlying network structures. However, many of the previous works have not answered a critical question: can the underlying model be identifiable with given data? It is possible that two different systems have the same dynamical profiles, meaning that the true model cannot be uniquely identified with given dynamical data. This indicates that the identifiability of the given system must be verified prior to the inference of a network structure and parameter estimation. In [9], I quantified the amount of transition rate information needed to uniquely identify the underlying biochemical network. I also proved that the time evolution of stochastic dynamics can be used to infer the hidden network structure, while information of deterministic dynamics may not uniquely characterize the network structure. We also proposed an algorithm that enables us to infer the underlying reaction network and the associated parameters with the transition data of a given continuous time Markov chain.
9. German Enciso, Radek Erban and Jinsu Kim, Identifiability of Stochastically Modelled Reaction Networks, accepted to European Journal of Applied Mathematics, 2020.
8. Jinsu Kim, Jason Dark, German Enciso and Suzanne Sindi, Slack Reactants: A State-Space Truncation Framework to Estimate Quantitative Behavior of the Chemical Master Equation, J. Chem. Phys. 153, 054117 (2020)
7. Enrico Bibbona, Jinsu Kim and Carsten Wiuf, Stationary distributions of systems with Discreteness Induced Transitions, J. R. Soc. Interface. 1720200243, 2020.
6. German Enciso and Jinsu Kim, Accuracy of Multiscale Reduction for Stochastic Reaction Systems, submitted, 2019.
5. David F. Anderson, Daniele Cappelletti, Jinsu Kim, and Tung Nguyen, Tier structure of strongly endotactic reaction networks, to appear in Stochastic Processes and their Applications .
4. David F. Anderson, Daniele Cappelletti and Jinsu Kim, Stochastically modeled weakly reversible reaction networks with a single linkage class, Journal of Applied Probability. 57(3), 792-810, 2020
3. Jinsu Kim and German Enciso , Absolutely Robust Controllers for Stochastic Chemical Reaction Networks, J. R. Soc. Interface. 17: 20200031, 2020.
2. German Enciso and Jinsu Kim, Embracing noise in chemical reaction networks, Bull Math Biol, 81, 1261–1267, 2019.
1. David F. Anderson and Jinsu Kim, Some network conditions for positive recurrence of stochastically modeled reaction networks, SIAM Journal of Applied Mathematics, 78(5), 2692-2713, 2018.
